|
8/22/2023 0 Comments Northern blot analysis biofilm Northern blot analysis of the hyphal wall protein (HWP1) expression was also monitored in planktonic and biofilm cells treated with EDTA. The extent and characteristics of biofilm formation were then microscopically assessed and with a semi-quantitative colorimetric technique based on the use of an XTT-reduction assay. All plates were then incubated at 37☌ for an additional 24 h to allow for biofilm formation. EDTA was also added to the standardized suspension prior to adding to the microtiter plate and to a preformed 24 h biofilm. Cells were allowed to adhere for 1, 2, and 4 h at 37☌, washed in PBS, and then treated with different concentrations of EDTA (0, 2.5, 25, and 250 mM). Candida albicans biofilms were formed in 96-well microtitre plates. In this study we investigated a means of inhibiting biofilm formation using EDTA (Ethylenediaminetetra-acetic acid), a divalent cation chelating agent, which has been shown to affect C. albicans biofilm etiology.Ībstract = "Candida albicans can readily form biofilms on both inanimate and biological surfaces. This compound may serve a non-toxic means of preventing biofilm formation on infections with a C. albicans biofilm formation are most likely through its inhibitory effect on filamentation and indicates the potential therapeutic effects of EDTA. 
These results indicate that EDTA inhibits C. The HWP1 gene expression was reduced in EDTA-treated planktonic and biofilm samples. However, preformed biofilms were minimally affected by EDTA (maximum of 31% reduction at 250 mM). Microscopic analysis and colorimetric readings revealed that filamentation and biofilm formation were inhibited by EDTA in a concentration dependant manner. Ensure that these reagents are in solution, and consider spinning in a microfuge or low speed centrifuge, or filtering the solutions through a 0.22 micron filter to remove particulates.Candida albicans can readily form biofilms on both inanimate and biological surfaces. Particulates in probe preparations or hybridization buffer (e.g., when not completely in solution) can also cause speckling on the membrane. Check probe quality and remove unincorporated nucleotides. Probe preparations with poor incorporation or where unincorporated nucleotides have not been removed, can cause speckling on the membrane. Use 10 pM nonisotopically labeled DNA probes and 0.1 nM nonisotopically labeled RNA probes. High probe concentrations, especially for nonisotopic probes, can also cause lane specific background. Start with a high hybridization temperature and slowly decrease temperature until specific signal is obtained. Hybridization conditions that are substantially below the optimum for a given probe can lead to high lane specific background and/or substantial cross-hybridization. 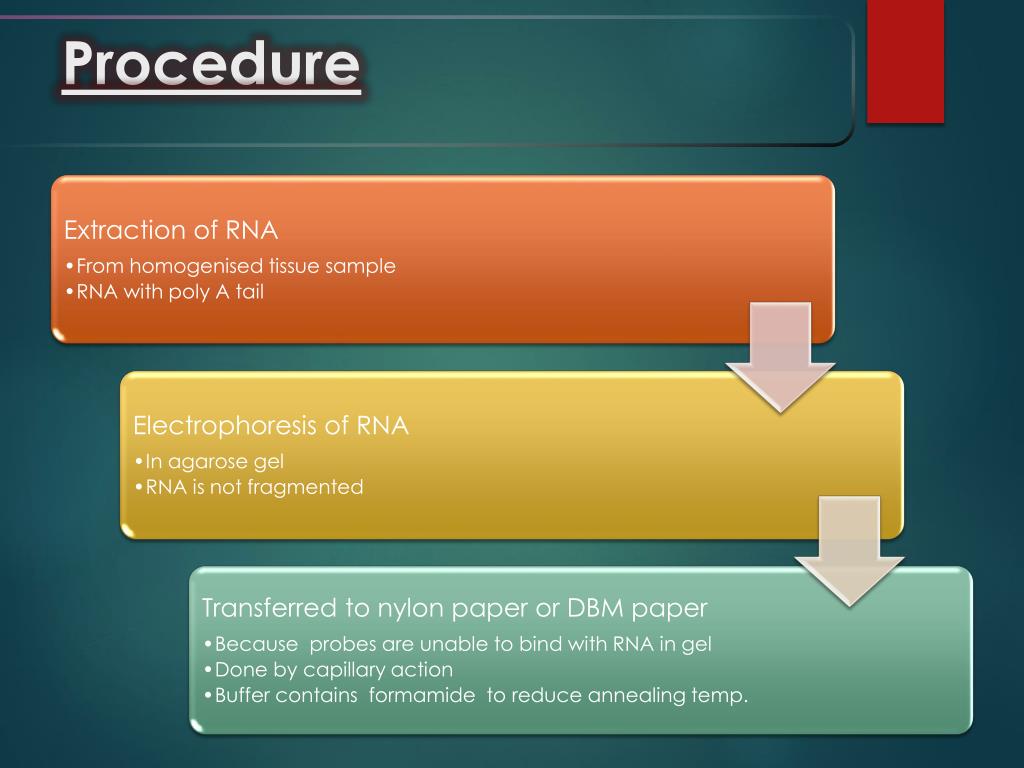
Do not pipet probe directly onto the membrane in hybridization solution dilute it into the hybridization solution first. Blotchiness can also be caused by uneven distribution of the hybridization reagents. Use high quality nylon membrane that has not previously been handled and use forceps to handle the membrane from the edges. Membrane of poor quality, that has dried out, or that has been mishandled (e.g., oil from human skin, powder from gloves) can cause this effect. There are several types of background, and each can have a different cause: Blotchy signal across the membrane 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
